|
mapgd
0.4
A program for the Maximum-likelihood analysis of population genomic data.
|
|
mapgd
0.4
A program for the Maximum-likelihood analysis of population genomic data.
|
#include <models.h>
Public Member Functions | |
| float_t | loglikelihood (const Locus &, const Allele &) |
| Gets the log likelihood of the observations at Locus, given Allele. More... | |
| float_t | goodness_of_fit (Locus &, const Allele &, std::vector< float_t > &, const float_t &) |
| Gets the log likelihood of the observations at Locus, given Allele. | |
| void | init_gof (const count_t *, const Allele &) |
| breif Initilizes the ? for goodness of fit calculations. | |
| float_t | get_gof (const count_t *, const Allele &) |
| Returns the likelihood of a ?? goodness of fit... blah blah blah. | |
| float_t | genotypelikelihood (const quartet_t &, const Allele &) |
| Clalculates genotypic likelihoods. Not implement, may be depricated. More... | |
| float_t | lnL (const float_t &logMM, const float_t &logMm, const float &logmm, const count_t *count) |
| Clalculates goodness of fit likelihoods. | |
| models & | operator= (const models &rhs) |
Private Attributes | |
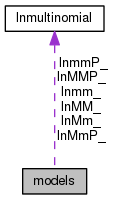
| lnmultinomial | lnMM_ |
| lnmultinomial | lnMm_ |
| lnmultinomial | lnmm_ |
| The three multinomials we will use for probability calculations. | |
| lnmultinomial | lnMMP_ |
| lnmultinomial | lnMmP_ |
| lnmultinomial | lnmmP_ |
| float_t | E0_ |
| float_t | E1_ |
| float_t | E2_ |
| Values used in calculations;. | |
this class perfoms some likelihood calculations. It keeps some look-up tables that will grow as calculations are performed, and may become to large over time. TODO: implement automatic look-up table purging.
Currently likelihood calculations are all performed by a set of 'models' that we give to the "multinomial" class via the multinomial::set method. These models should be able to look at the Allele structure, which contains information about the error rate at the locus and the identity of the major and minor allele, and return a set of four log probabilities of observing each particular nucleotide in a given call.
Clalculates genotypic likelihoods. Not implement, may be depricated.
DONT USE THIS!!! TODO FIX IT!.
 1.8.6
1.8.6