|
|
Allele & | operator= (const Allele &) |
| | use the = operator to assign Allele.
|
| |
|
| Allele (std::vector< std::string >) |
| | Instance of constructor required by Data, but Allele doesn't actually need the vector to initialize.
|
| |
|
std::string | header (void) const |
| | returns first two lines of a file.
|
| |
|
size_t | size (void) const |
| | The size of the class in bytes.
|
| |
|
const std::string | get_file_name (void) const |
| | Returns the default file extension.
|
| |
|
const std::string | get_table_name (void) const |
| | Returns the destination table name.
|
| |
|
const bool | get_binary (void) const |
| | Returns the destination table name.
|
| |
|
const std::string | sql_header (void) const |
| | Returns a string to create an SQL table.
|
| |
|
const std::string | sql_column_names (void) const |
| | Returns the column names in the SQL table.
|
| |
|
const std::string | sql_values (void) const |
| | Returns a string to insert values in an SQL table.
|
| |
|
void | write_pos (std::ostream &str) const |
| |
|
void | read_pos (std::istream &str) |
| |
|
| Indexed_data (std::vector< std::string > &) |
| |
|
id1_t | get_abs_pos (void) const |
| | Indexed data needs to associate each datum with a position in the genome.
|
| |
|
void | set_abs_pos (const id1_t &) |
| | Indexed data needs to associate each datum with a position in the genome.
|
| |
|
const bool | indexed () const |
| |
|
void | read_binary (std::istream &str) |
| |
|
void | write_binary (std::ostream &str) const |
| |
|
| Data (std::vector< std::string > &) |
| |
|
virtual void | sql_read (std::istream &) |
| | Reads the values...
|
| |
|
|
char | delim |
| | the delimiter used when reading/writing the class in text mode.
|
| |
|
count_t | excluded |
| | A count of the number of samples that were excluded due to filtering criteria.
|
| |
|
bool | pooled |
| | Inferred from pooled or labeled sequencing?
|
| |
|
float_t | freq |
| | frequency of major allele.
|
| |
|
gt_t | ref |
| | identity of ref allele.
|
| |
|
gt_t | minor |
| | identity of minor allele.
|
| |
|
gt_t | major |
| | identity major allele.
|
| |
|
gt_t | e1 |
| | identity of error1
|
| |
|
gt_t | e2 |
| | identity of error2.
|
| |
|
float_t | error |
| | ml error rate.
|
| |
|
float_t | null_error |
| | error rate assuming Major allele monomorphism.
|
| |
|
float_t | null_error2 |
| | error rate assuming Minor allele monomorphism.
|
| |
|
count_t | coverage |
| | population coverage.
|
| |
|
float_t | efc |
| | number of 'effective' chromosomes in the sample.
|
| |
|
float_t | ll |
| |
|
float_t | monoll |
| |
|
float_t | hwell |
| | log likelihoods.
|
| |
|
float_t | MM |
| | frequency of genotype MM in the population.
|
| |
|
float_t | Mm |
| | frequency of genotype Mm in the population.
|
| |
|
float_t | mm |
| | frequency of genotype mm in the population.
|
| |
|
count_t | N |
| | number of individual at the site.
|
| |
|
float_t | f |
| | HW statistic.
|
| |
|
float_t | h |
| | heterozygosity.
|
| |
|
float_t | gof |
| | gof statistic.
|
| |
Summary statistics from the allele command.
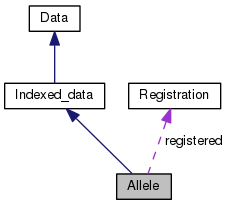
 Public Member Functions inherited from Indexed_data
Public Member Functions inherited from Indexed_data Public Member Functions inherited from Data
Public Member Functions inherited from Data Static Public Attributes inherited from Data
Static Public Attributes inherited from Data Static Public Member Functions inherited from Data
Static Public Member Functions inherited from Data Protected Attributes inherited from Indexed_data
Protected Attributes inherited from Indexed_data 1.8.6
1.8.6