|
mapgd
0.4
A program for the Maximum-likelihood analysis of population genomic data.
|
|
mapgd
0.4
A program for the Maximum-likelihood analysis of population genomic data.
|
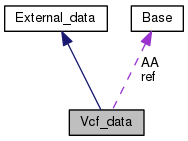
Public Member Functions | |
| Vcf_data () | |
| number of individuals sapmled | |
| void | set_header (const File_index &, const std::vector< std::string > &) |
| void | put (const Data *,...) |
| The write function must be defined in the child class. | |
| void | get (Data *,...) const |
| The read function must be defined in the child class. | |
| void | put (const File_index &, const Allele &, const Population &) |
| id1_t | get (const File_index &, Population &) const |
| id1_t | get (State &) const |
| std::vector< std::string > | get_sample_names (void) const |
| File_index | get_index (void) const |
Data Fields | |
| bcf1_t * | record_ |
| bcf_hdr_t * | header_ |
| size_t | sample_size_ |
| std::string | id |
| Base | ref |
| std::vector< std::string > | alt |
| float_t | qual |
| bool | filter |
| std::vector< std::string > | info |
| Base | AA |
| Ancestral allele. | |
| std::vector< count_t > | AC |
| Allele count in genotypes, for each ALT allele, in the same order as listed. | |
| std::vector< float_t > | AF |
| Allele frequency for each ALT allele in the same order as listed: | |
| count_t | AN |
| Total number of alleles in called genotypes. | |
| float_t | BQ |
| RMS base quality at this position. | |
| bool | DB |
| Cigar string describing how to align an alternate allele to the reference allele. More... | |
| count_t | DP |
| Combined depth across samples, e.g. DP=154. | |
| id1_t | END |
| End position of the variant described in this record (esp. for CNVs) | |
| bool | H2 |
| Membership in hapmap2. | |
| float_t | MQ |
| RMS mapping quality, e.g. MQ=52. | |
| count_t | MQ0 |
| Number of MAPQ == 0 reads covering this record. | |
| count_t | NS |
| Number of samples with data. | |
| float_t | SB |
| Strand bias at this position. | |
| bool | SOMATIC |
| Indicates that the record is a somatic mutation, for cancer genomics. | |
| bool | VALIDATED |
| Validated by follow-up experiment. | |
| bool Vcf_data::DB |
Cigar string describing how to align an alternate allele to the reference allele.
dbSNP membership
 1.8.6
1.8.6