|
mapgd
0.4
A program for the Maximum-likelihood analysis of population genomic data.
|
|
mapgd
0.4
A program for the Maximum-likelihood analysis of population genomic data.
|
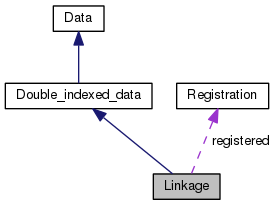
A class to store linkage information. More...
#include <linkage_data.h>
Public Member Functions | |
| Linkage (const std::vector< std::string > &) | |
| std::string | header (void) const |
| size_t | size (void) const |
| The size of the class in bytes. | |
| const std::string | get_file_name (void) const |
| The default extension for files. | |
| const std::string | get_table_name (void) const |
| The default extension for files. | |
| const bool | get_binary (void) const |
| The default extension for files. | |
| const std::string | sql_header (void) const |
| Destination table in Db. | |
| const std::string | sql_column_names (void) const |
| Destination table in Db. | |
| const std::string | sql_values (void) const |
| Destination table in Db. | |
| void | set_D (const float_t &) |
| void | set_fit (const float_t &) |
| void | set_null (const float_t &) |
| void | set_Ni (const float_t &) |
| void | set_p (const float_t &) |
| void | set_q (const float_t &) |
| float_t | ll (void) const |
| float_t | dist (void) const |
| float_t | D (void) const |
| float_t | Dmax (void) const |
| float_t | Dmin (void) const |
| float_t | Dsq (void) const |
| float_t | Dprime (void) const |
| float_t | r (void) const |
| float_t | rsq (void) const |
| float_t | adj_D (void) const |
| float_t | adj_Dsq (void) const |
| float_t | adj_Dprime (void) const |
| float_t | adj_r (void) const |
| float_t | adj_rsq (void) const |
| float_t | adj_Dmax (void) const |
| float_t | adj_Dmin (void) const |
| Linkage & | operator= (const Linkage &lhs) |
 Public Member Functions inherited from Double_indexed_data Public Member Functions inherited from Double_indexed_data | |
| void | write_pos (std::ostream &str) const |
| void | read_pos (std::istream &str) |
| Double_indexed_data (std::vector< std::string > &) | |
| id1_t | get_abs_pos (void) const |
| Indexed data needs to associate each datum with a position in the genome. | |
| id1_t | get_abs_pos2 (void) const |
| Indexed data needs to associate each datum with a position in the genome. | |
| void | set_abs_pos (const id1_t &) |
| Indexed data needs to associate each datum with a position in the genome. | |
| void | set_abs_pos2 (const id1_t &) |
| Indexed data needs to associate each datum with a position in the genome. | |
| const bool | indexed () const |
 Public Member Functions inherited from Data Public Member Functions inherited from Data | |
| void | read_binary (std::istream &str) |
| void | write_binary (std::ostream &str) const |
| Data (std::vector< std::string > &) | |
| virtual void | sql_read (std::istream &) |
| Reads the values... | |
Data Fields | |
| char | delim |
| < Set the absolute position of the Locus. More... | |
Static Public Attributes | |
| static const std::string | file_name =".lds" |
| delegating the constructor ftw. More... | |
| static const std::string | table_name ="LINKAGE" |
| Destination table in Db. | |
| static const bool | binary =false |
| Returns the destination table name. | |
 Static Public Attributes inherited from Data Static Public Attributes inherited from Data | |
| static const std::string | file_name =".txt" |
| The read function must be defined in the child class. More... | |
| static const std::string | table_name ="NONE" |
| The read function must be defined in the child class. More... | |
Private Member Functions | |
| void | read (std::istream &str) |
| The read function must be defined in the child class. | |
| void | write (std::ostream &str) const |
| ! the write function must be ? by the child class. | |
Static Private Member Functions | |
| static Data * | create (const std::vector< std::string > &columns) |
Private Attributes | |
| id0_t | id0 |
| The scaffold of site X. | |
| id1_t | id1 |
| The pos of site X. | |
| float_t | p_ |
| freq_major site numero uno | |
| float_t | q_ |
| freq_major site numero dos | |
| float_t | D_ |
| the magnitude of the ld, j0 | |
| float_t | Ni_ |
| The number of individuals used in the calculation. | |
| float_t | fit_ |
| fit statistic (thar be ld here!) | |
| float_t | null_ |
| null statistic (i.e. no ld) | |
Static Private Attributes | |
| static const Registration | registered =Registration(Linkage::table_name, Linkage::create) |
Additional Inherited Members | |
 Static Public Member Functions inherited from Data Static Public Member Functions inherited from Data | |
| static Data * | new_from_str (const std::string &, const std::vector< std::string > &) |
| Constructs an instance of the class Registered w/ string. | |
 Protected Attributes inherited from Double_indexed_data Protected Attributes inherited from Double_indexed_data | |
| id1_t | abs_pos1_ |
| id1_t | abs_pos2_ |
A class to store linkage information.
Many of these data classes need to have things moved over to static functions. The problem popped up because I just copied over all the output people wanted into values in the class, but really a lot of this is just simple transformation of other representions of the data. If you are reading this may god have mercy on your soul, because I really have no idea why you would choose to read the comments on what is doubtlessly a dead project. please enjoy this link to a Kraftwork video to see if Doxygen automatically incorporates HTML tags right (IT DOES!). Listening to Kraftwerk improves your coding skillz. It's totally a fact. I'll start fixing allele stat now.
< The scaffold of site X
< The pos of site X
< freq_major site numero uno
< freq_major site numero dos
< the magnitude of the ld, j0
< The number of individuals used in the calculation
< fit statistic (thar be ld here!)
< null statistic (i.e. no ld)
| char Linkage::delim |
< Set the absolute position of the Locus.
the delimiter used when reading/writing the class in text mode.
|
static |
delegating the constructor ftw.
The default extension for files.
 1.8.6
1.8.6